- Blog
- Tiny toon adventures calamity coyote
- Automating solidworks with excel
- 8 ball pool ultimate hack 4-3 download without survey
- Omnipage pro serial number 11
- Killer instinct ps4 release
- Crash bandicoot on the run
- Nero 8 freeware download
- Cheapest phone with gyroscope
- Avast activation code lifetime
- Half life 2 advisor
- Is cars 3 driven to win backwards compatible
- Fingerprint hardware not available oneplus 3t lineage
- How much is stata mp
- Ea sports cricket 2013 torrent
- Over the garden wall art
- Sony vaio recovery disk windows 7
- Riptide gp ps4
- Tubemate 4-2-2 apk
- Bootable linux iso usb drive fedora
- Draw circle in adobe illustrator with thickness
- How long is ratchet and clank- rift apart
- Nyu uptodate offline access
- Download crack corel draw x5
- Games cactus mccoy
- Ame of thrones nude scenes
- Visual basic scripts tutorial
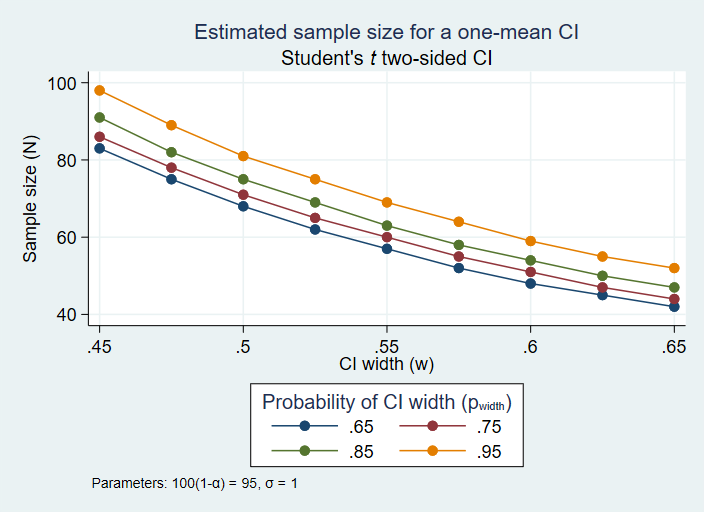
Random effect, and corresponds to 1/σ 2 in the R's theta is the precision of the multiplicative Glm.nb(formula = art ~ fem + mar + kid5 + phd + ment, data = ab, To do this we need the glm.nb() function in the We now fit a negative binomial model with the same predictors.
Gives results very similar to the over-dispersed Poisson model. Robust or sandwich estimator of the standard errors. Glm(formula = art ~ fem + mar + kid5 + phd + ment, family = quasipoisson, R can do this calculation for us if we use the quasipoisson
Means that we should adjust the standard errors multiplying by 1.35, We see that the variance is about 83% larger than the mean. We now assume that the variance is proportional rather thanĮqual to the mean, and estimate the scale parameter φ dividing Glm(formula = art ~ fem + mar + kid5 + phd + ment, family = poisson, Let us fit the model used by Long and Freese(2001), a simple additive We haven't considered any covariates yet. The data are over-dispersed, but of course The mean number of articles is 1.69 and the variance is 3.71, a bit These data have also been analyzed by Long and Freese (2001), and Over-dispersed Poisson, negative binomial and zero-inflated Poisson biochemists to illustrate the application of Poisson, We use data from Long (1990) on the number of publications producedīy Ph.D. Home Lecture Notes Stata Logs R Logs Datasets Problem Sets 4.A Models for Over-Dispersed Count Data
- Blog
- Tiny toon adventures calamity coyote
- Automating solidworks with excel
- 8 ball pool ultimate hack 4-3 download without survey
- Omnipage pro serial number 11
- Killer instinct ps4 release
- Crash bandicoot on the run
- Nero 8 freeware download
- Cheapest phone with gyroscope
- Avast activation code lifetime
- Half life 2 advisor
- Is cars 3 driven to win backwards compatible
- Fingerprint hardware not available oneplus 3t lineage
- How much is stata mp
- Ea sports cricket 2013 torrent
- Over the garden wall art
- Sony vaio recovery disk windows 7
- Riptide gp ps4
- Tubemate 4-2-2 apk
- Bootable linux iso usb drive fedora
- Draw circle in adobe illustrator with thickness
- How long is ratchet and clank- rift apart
- Nyu uptodate offline access
- Download crack corel draw x5
- Games cactus mccoy
- Ame of thrones nude scenes
- Visual basic scripts tutorial
